
Sequence Braiding
Sara Di Bartolomeo, Yixuan Zhang, Fangfang Sheng, Cody Dunne
Preprint on OSF: https://osf.io/mq2wt/Repository on github: https://github.com/VisDunneRight/sequence_braiding
Supplemental material (code, data, results): https://osf.io/s92bu/
Temporal event sequence alignment has been used in many domains to visualize nuanced changes and interactions over time. Existing approaches align one or two sentinel events. Overview tasks require examining all alignments of interest using interaction and time or juxtaposition of many visualizations. Furthermore, any event attribute overviews are not closely tied to sequence visualizations. We present Sequence Braiding, a novel overview visualization for temporal event sequences and attributes using a layered directed acyclic network. Sequence Braiding visually aligns many temporal events and attribute groups simultaneously and supports arbitrary ordering, absence, and duplication of events. In a controlled experiment we compare Sequence Braiding and IDMVis on user task completion time, correctness, error, and confidence. Our results provide good evidence that users of Sequence Braiding can understand high-level patterns and trends faster and with similar error.
Table of Contents
Dataset format
Sample dataset for three days:Try editing me![
[
{"type": "Breakfast", "level": "very_high"},
{"type": "Lunch", "level": "very_high"},
{"type": "Snack", "level": "very_high"},
{"type": "Dinner", "level": "very_high"}
],
[
{"type": "Breakfast", "level": "very_high"},
{"type": "Snack", "level": "high"},
{"type": "Dinner", "level": "very_high"}
],
[
{"type": "Breakfast", "level": "high"},
{"type": "Lunch", "level": "very_high"},
{"type": "Snack", "level": "high"},
{"type": "Dinner", "level": "high"}
]
]
Playground
Case Studies and examples
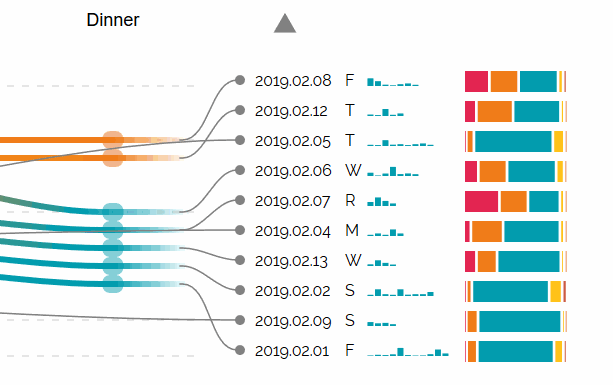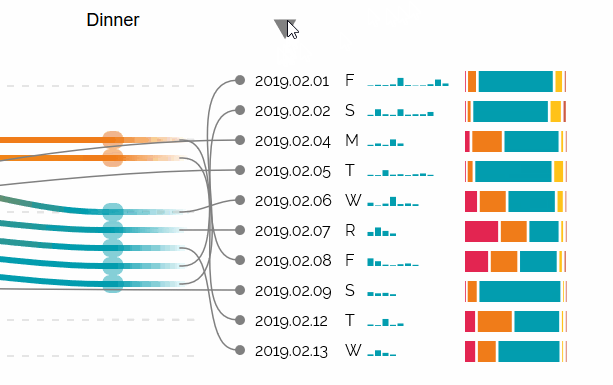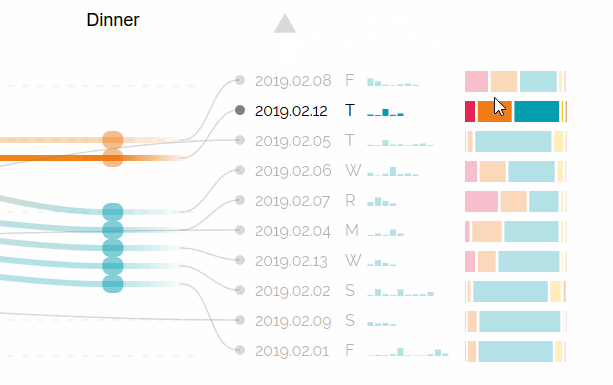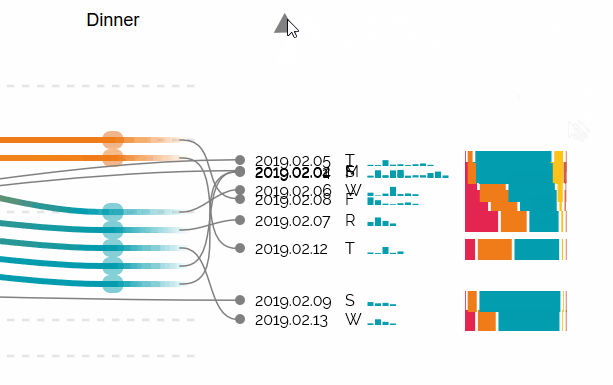
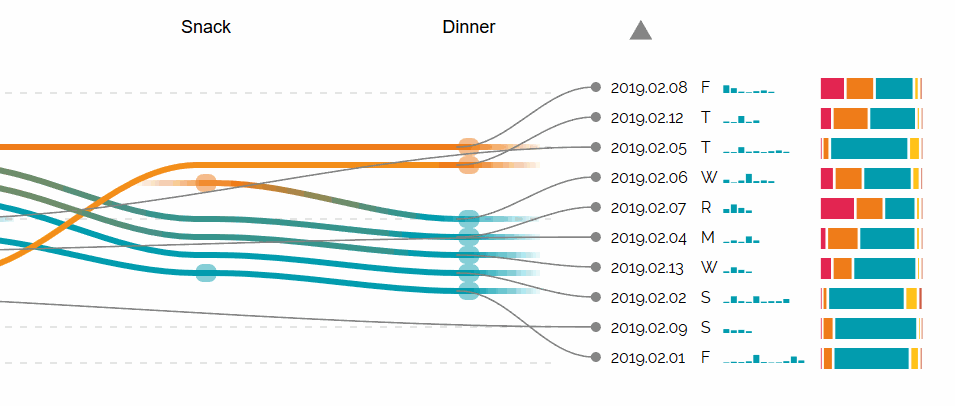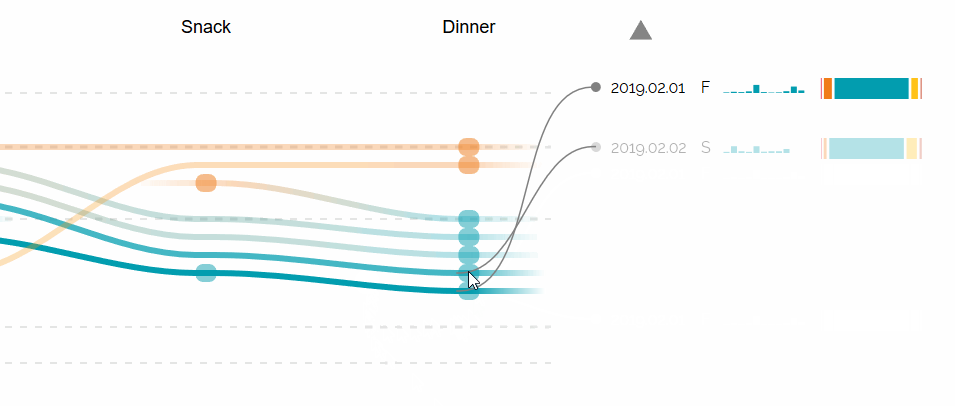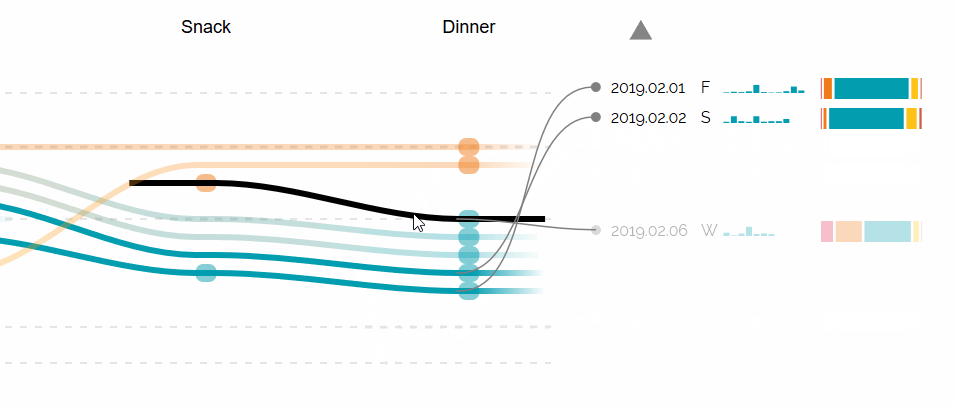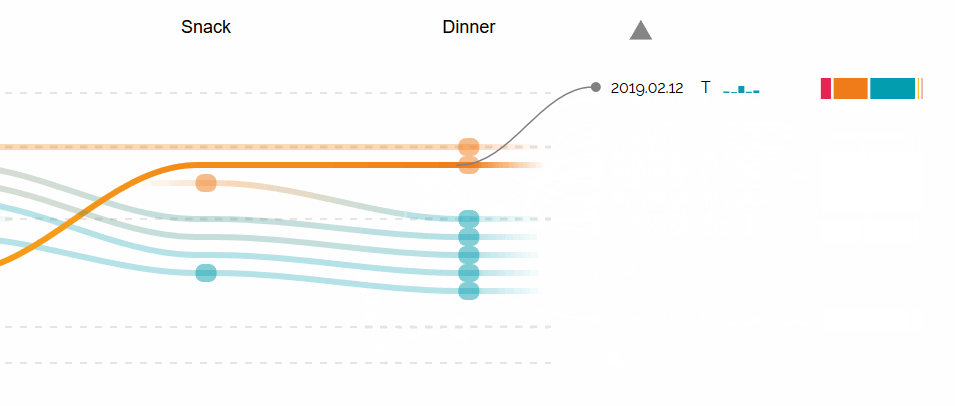
Augmented Sequence Braiding, a plugin to display additional features associated to the sequences.
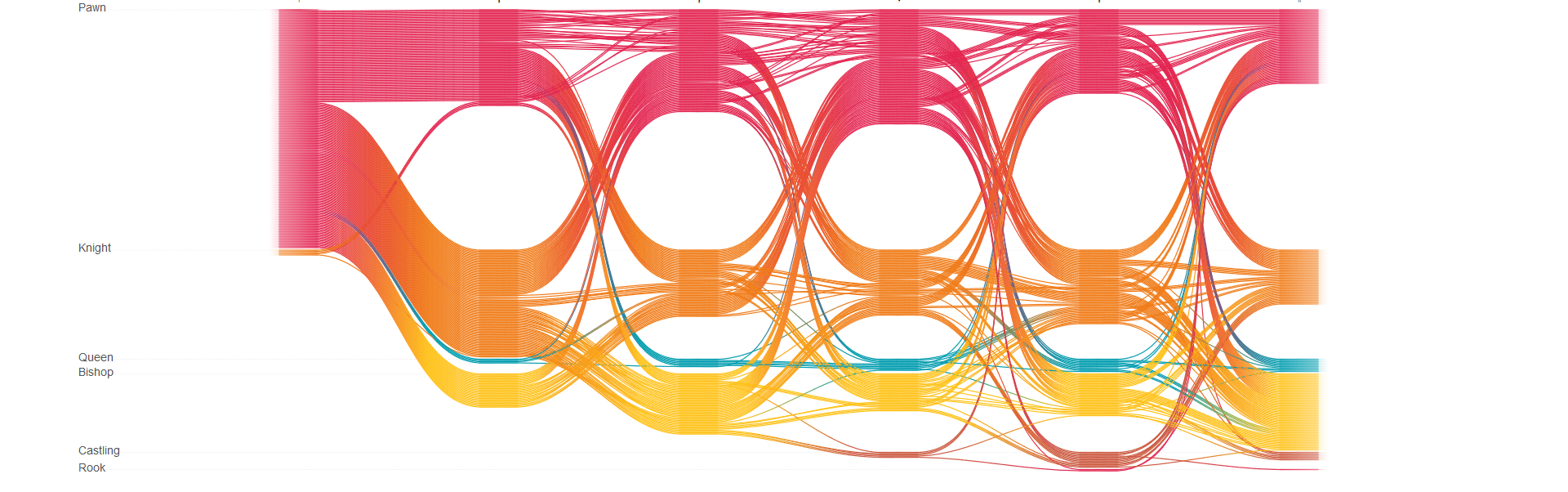

Chess, a visualization of the sequences of moves played in 200 games of chess.
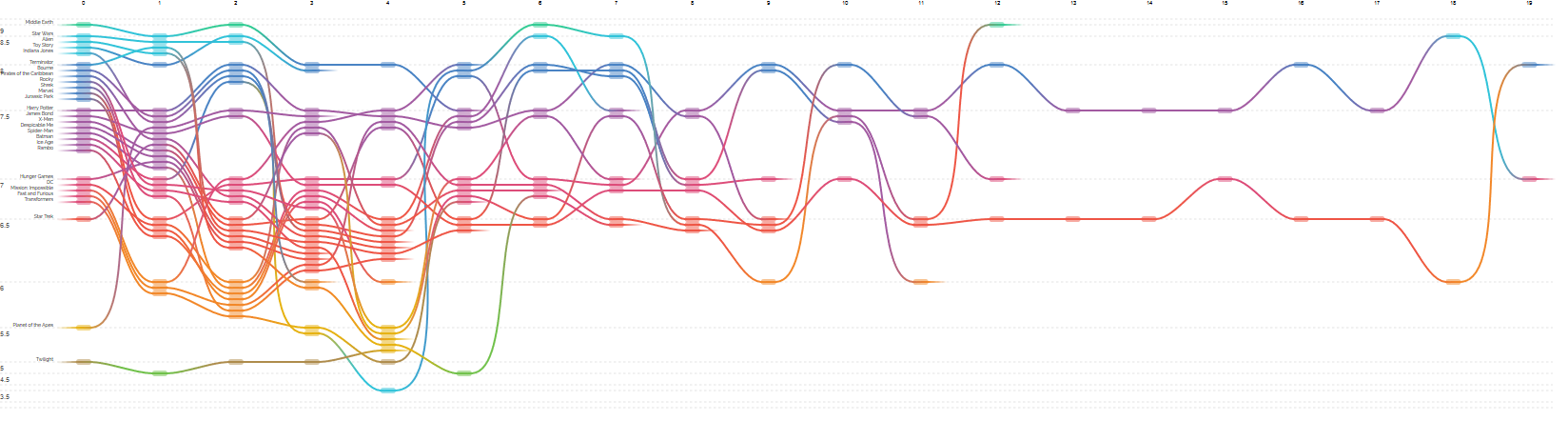
How many movies does it take to kill a franchise?, a visualization of the IMDb ratings of sequences of movies in the same franchise.
Usage
Initialize the graph with: var graph = new SequenceBraiding(data, "svgname", options)svgname: the id of the svg in which you want to draw the graph.
options: an object containing the parameters for drawing the graph.
Options
The options object is a javascript object that allows you to customize Sequence Braiding in terms of visual representation or to give parameters to the algorithm.Example:
graph_options = {
numSequences: 10,
minEventPerColThreshold: 1,
height: 1000,
width: window.innerWidth*0.7,
forceLevelName: true,
node_width_factor: 0.08,
levels: ['very_high', 'high', 'normal', 'low', 'very_low'],
}Features
guidelines: draws faint lines between a level and the other to help distinguish them.Default: true
animate: activates or deactivates animations in the graph, which show how the nodes are moved at each iteration of the intersection reduction algorithm.
Default: false
colorbysequence: if set to true, each sequence line will have a specific color, and the colors won't change according to the node level, but will be the same color throughout the full sequence.
Default: false
forceLevelName: forces the level name to be displayed even with a high number of lines.
Default: false
Data parameters
numSequences: determines the maximum amount of sequences to display.Default: 10
minEventPerColThreshold: columns that contain a number of events under this threshold will be filtered out.
Default: 1
Algorithm parameters
max_iterations: maximum number of iterations to be run by the intersection reduction algorithm.Default: 20
path: can be used to force the order of events. If left undefined, the algorithm will guess it while also trying to reduce the intersections as much as possible.
Example: path: ["source", "breakfast", "lunch", "sink"]
Default: undefined
levels: can be used to force the order of levels. Especially useful when the levels have a sequential ordering. If left undefined, the algorithm will try to guess it.
Default: undefined
Pairwise alignment parameters
Pairwise alignment parameters are used to generate the minimum supersequence of event types. If path is not forced, Sequence Braiding will use these parameters to generate a supersequence using pairwise DNA alignment. Depending on the needs of the specific user, these parameters can be changed. pairwise_alignment_vars must be given as an input as a subfield of the options object.options = {
pairwise_alignment_vars: {
MATCH_SCORE: 100,
MISMATCH_SCORE: -10,
BEGIN_GAP_PENALTY: 2,
GAP_PENALTY: 1,
END_GAP_PENALTY: 2,
},
//...
}Graphical presentation
height: height of the visualization.Default: 400
width: width of the visualization.
Default: '100%'
svg_width: width of the svg.
Default: '100%'
link_opacity: opacity of the lines.
Default: 1
node_width_factor: size of the nodes.
Default: 0.2
link_stroke_width: size of the lines.
Default: 4
padding: takes an object that describes the top and the left padding of the visualization - not of the text.
Default: {top: 40, left: 40}
colorscheme: takes an array of strings of hex values of colors. Colors used for the visualization.
Default: ["#E32551", "#F07C19", "#029DAF", "#FFC219", "#cd5b43"]
fontSize: size of the fonts used in the graph.
Default: '0.9em'
catmullromvalue: how smooth the curves appear. Closer to 0 will produce more rough curves, but will be more readable on more cluttered visualizations. Closer to 1 will produce softer curves.
Default: 1
path_text_y: the vertical position of the text containing the event types (at the top of each column).
Default: 10